neuroelf_gui_-_menu_tree
This is an old revision of the document!
Table of Contents
NeuroElf - GUI menu tree
Motivation
This page is intended to give an overview on functions and settings available via the main menu. Each item is listed in the order of appearance and a brief explanation is given of what function is performed or how the settings are changed.
Requirements
To follow this page, you need to have the GUI open by entering neuroelf_gui at the Matlab prompt.
Background information
Configuration settings that persist across NeuroElf GUI sessions are stored in a text-based configuration (ini) file in _core/config/neuroelf.ini. Some of these settings can be reset to their original state by issuing neuroelf_setup.
File menu
- Open… - opens a file selector for all major formats available for visualization, both anatomical/functional (VMR, MSK, HDR/NII, HEAD, VTC, FMR, DMR,and also SRF) as well as statistical (VMP, GLM, HDR/NII, HEAD, CMP/ICA, as well as SMP) will be listed by default; if the filetype selector (type restriction on displayed items) is changed, this selection will be remembered, additionally NeuroElf changes the current working directory (Matlab's
pwd) to the folder that contains the file that is being opened - Select current object - a sub-menu populated with all loaded (voxel-based) anatomical objects (only visible when the GUI is in minimized mode)
- Recently loaded slicing objects - a sub-menu populated with anatomical (voxel-based) files that were recently opened (only if opened via the GUI and file selection dialog, not via a script or
object.Browse;call; this information is stored in the configuration file under section [RecentFiles], settingSliceVar) - Recently loaded slicing stats objects - a sub-menu populated with statistical (voxel-based) files that were recently opened (stored in setting
StatsVar) - Recently loaded surface objects - a sub-menu populated with surface files that were recently opened (stored in setting
SurfVar) - Recently loaded surface stats objects - a sub-menu populated with surface-based statistics files that were recently opened (stored in setting
SurfStatsVar) - Colin-27 dataset - a sub-menu populated with the files found in the
_files/colin/' sub-folder of the NeuroElf installation folder (in two sub-menus: VMRs and Surfaces) * Clone slicing object - creates a copy of the currently selected slicing (voxel-based anatomical or functional) object, e.g. to allow painting into the object with the ability to compare “before” and “after” stages; this does not copy the filename association (and thus, the object requires to be “saved as…”) * Reload slicing object - reloads the content of the currently selected slicing object from disk (only functional if the object has been loaded or saved before and a filename associated with it) * Reload stats object - reloads the content of the currently selected (voxel-based) statistical object from disk (same restriction applies) * Set underlay object - sets a secondary object which is display below (underlaying) the current selected anatomical slicing object (the way in which the two datasets are blended can be controlled via the File → Options → Underlay blending mode sub-menu) * Save - saves (i.e. overwrites) the currently selected slicing (voxel-based anatomical or functional) object back to disk (only functional if the object has been loaded or saved before and a filename associated with it) * Save as… - opens a file saving dialog (uiputfile), and, unless this dialog is canceled or an invalid filename is entered, stores the currently selected slicing object under the given filename (and stores this filename with the object for subsequent calls toFile→Save) * Save stats - saves (i.e. overwrites) the currently selected (voxel-based) statistical object back to disk (only functional if the object has been loaded or saved before and a filename associated with it) * Save stats as… - opens a file saving dialog (uiputfile), and, unless this dialog is canceled or an invalid filename is entered, stores the currently selected (voxel-based) statistical object under the given filename (and stores this filename with the object for subsequent calls toFile→Save) * Import Analyze to VMR… - allows to select an Analyze/NIftI image, which is then converted to VMR format using the importvmrfromanalyze function and subsequently loaded and displayed in the GUI * Import SPM maps to VMP… - allows to select a set of SPM-based statistic files (spm_Fxxxx.hdr/niiand/orspm_Txxxx.hdr/niifiles) which are then imported to VMP format using the importvmpfromspms function and subsequently loaded and displayed in the GUI * Import RFX-GLM from SPM.mat files… - opens the importrfxglmfromspms UI to import a RFX-GLM file from SPM.mat files ==== File → Options menu ==== * ClusterTable - contains settings changing the cluster table generation feature (behavior when using theClustertablebutton) * generate new VOI - if selected (and subsequently being “checked”; this is the default) each use of theClustertablebutton will remove all currently available clusters (VOIs) from the GUI before the clusters of the currently selected map will be stored * add to existing VOI - if selected (and subsequently being “checked”) each use of theClustertablebutton will add clusters as found by the VMP::ClusterTable method to the list of available clusters * peak coordinate - if selected (and subsequently being “checked”; this is the default) the text output (table of clusters) will show the peak coordinate in each cluster (highest absolute value of statistics for a consecutive, coherent set of coordinates) for each entry * center coordinate - if selected (and subsequently being “checked”) the text output will show the average (mean) coordinate for each cluster * center-of-gravity - if selected (and subsequently being “checked”) the text output will show a weighted average (sum of the products of absolute statistical values and coordinates divided by the sum of the absolute statistical values)Renderer
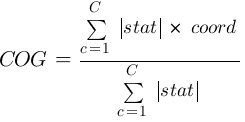 * print sub-cluster sizes - if selected (and subsequently being “checked”) the text output will contain an estimate of the number of voxels also for sub-clusters (using a watershed-based algorithm)
* sort table by stats - if selected (and subsequently being “checked”) the list of clusters will be sorted by the (absolute) stats value (decending values)
* sort table by signed stats - if selected (and subsequently being “checked”; this is the default) the list of clusters will first be sorted by the sign of each cluster peak and then, within positive and negative clusters, by the absolute value of the peak coordinate (descending values)
* sort table by cluster size - if selected (and subsequently being “checked”) the list of clusters will be sorted by the cluster size of each cluster (decending values)
* sort table by X coordinate - if selected (and subsequently being “checked”) the list of clusters will be sorted by the X coordinate of each cluster peak (decending values)
* sort table by Y coordinate - if selected (and subsequently being “checked”) the list of clusters will be sorted by the Y coordinate of each cluster peak (decending values)
* sort table by Z coordinate - if selected (and subsequently being “checked”) the list of clusters will be sorted by the Z coordinate of each cluster peak (decending values)
* Orientation - sub-item menu with…
* neurological - if selected (and subsequently being “checked”; this is the default) voxel-based datasets will be shown with the left side of axial and coronal images representing the left hemisphere (left-is-left)
* radiological - if selected (and subsequently being “checked”) datasets will be shown with the left side of axial and coronal images representing the right hemisphere (left-is-right)
* Renderer - sub-item menu with…
* OpenGL - if selected (and subsequently being “checked”; this is the default) the main UI figure (as well as satellite figures) will have their
* print sub-cluster sizes - if selected (and subsequently being “checked”) the text output will contain an estimate of the number of voxels also for sub-clusters (using a watershed-based algorithm)
* sort table by stats - if selected (and subsequently being “checked”) the list of clusters will be sorted by the (absolute) stats value (decending values)
* sort table by signed stats - if selected (and subsequently being “checked”; this is the default) the list of clusters will first be sorted by the sign of each cluster peak and then, within positive and negative clusters, by the absolute value of the peak coordinate (descending values)
* sort table by cluster size - if selected (and subsequently being “checked”) the list of clusters will be sorted by the cluster size of each cluster (decending values)
* sort table by X coordinate - if selected (and subsequently being “checked”) the list of clusters will be sorted by the X coordinate of each cluster peak (decending values)
* sort table by Y coordinate - if selected (and subsequently being “checked”) the list of clusters will be sorted by the Y coordinate of each cluster peak (decending values)
* sort table by Z coordinate - if selected (and subsequently being “checked”) the list of clusters will be sorted by the Z coordinate of each cluster peak (decending values)
* Orientation - sub-item menu with…
* neurological - if selected (and subsequently being “checked”; this is the default) voxel-based datasets will be shown with the left side of axial and coronal images representing the left hemisphere (left-is-left)
* radiological - if selected (and subsequently being “checked”) datasets will be shown with the left side of axial and coronal images representing the right hemisphere (left-is-right)
* Renderer - sub-item menu with…
* OpenGL - if selected (and subsequently being “checked”; this is the default) the main UI figure (as well as satellite figures) will have their property set to'OpenGL' (using hardware-based acceleration when possible) * zbuffer - if selected (and subsequently being “checked”) the figures will have theirRendererproperty set to'zbuffer' * Stats Colors (LUT) - a sub-menu populated with the OLT (overlay look-up table) files available in the_core/lutfolder of the NeuroElf installation; for each of the sub-entries, the currently selectedMap(only functional if exactly one map is selected) will have itsLUTNameproperty set, followed by a call toobject.SetColors, which will update the coloring of the map if, and only if, the map'sUseRGBColorproperty is set to 0 (changeable by clicking on theLUTradio button on the interface) * Edit LUT palette - allows to edit the currently selected LUT palette using the colorpicker function/UI (if 20 colors, used for VMR coloring) * *Underlay blending mode - a sub-menu with 7 different settings on how the underlay object (if selected) is blended with the currently selected anatomical slicing object * Overlay wins - wherever the overlay (selected object) has a voxel color that is not all-0, this value will be display, the underlay value otherwise * Overlay favored - the higher the maximum RGB color value of the overlay (select object) voxel is, the more this object's color dominates the mix * Overlay weighted - the ratio of voxel colors is determined byvalue_overlay / (value_overlay + value_underlay)* Equal mixing - voxel color values are mixed by a 50:50 ratio * Underlay weighted - the ratio of voxel colors is determined byvalue_underlay / (value_overlay + value_underlay)* Underlay favored - the higher the maximum RGB color value of the underlay object voxel is, the more this object's color dominates the mix * Underlay wins - wherever the underlay object has a voxel color that is not all-0, this value will be display, the overlay (selected object) value otherwise * Linked browsing with satellites - if checked (selection of this entry checks and un-checks this item) browsing (clicking into) a voxel-based dataset will set the cursor position (current coordinate) not only in the window that is clicked into but in all available windows (main UI and satellites) * Show stats thresh bars - if checked (selection of this entry checks and un-checks this item) a colored bar will be shown for each selected statistical map (only selected tails; and currently only implemented for voxel-based datasets, whereas if three panels are shown the bars will be on the coronal slice and if only a single panel is shown the bars will be doubled in size) * Echo function call/methods - if checked (selection of this entry checks and un-checks this item) several calls (menu items, button clicks, etc.) will result in a line of code being printed to the Matlab command window which can be used as a sort of template to write scripts controlling UI behavior ===== File menu (continued…) ===== * Exit - this closes the main UI, after a request dialog is OK-ed; if desired, all objects that were loaded by the UI (and not, for instance, passed into the UI viaobject.Browse;calls!), will be deleted from the xff global object memory ===== VMR menu ===== This menu is only visible if the currently selected slicing (voxel-based anatomical) object is of type VMR. * Border reconstruction - the VMR::DBReco method will be called on the currently selected VMR object, resulting in a SRF (surface) object that will subsequently be added to the NeuroElf GUI interface; this function is only useful if the currently selected VMR has been “segmented” (voxels above a certain threshold represent white matter voxels)! * Clean VMR - call the VMR::CleanVMR method to create a second (copy) of the VMR with all voxels detected as background set to 0 * Correct inhomogeneity - call the VMR::InhomogeneityCorrect method, which in turn uses the pmbfilter function in 3 passes to auto-correct the inhomogeneity of the data * Smooth VMR - this opens up a small dialog box, where smoothing kernel, bounding box, as well as an optional value range can be specified; subsequently the AFT::SmoothData3D method will be called to perform the data smoothing (for now, this may require a large amount of memory, depending on the settings; this might lead to out-of-memory error conditions, particularly for 0.5mm-resolution VMRs!) * Load Undo-Buffer… - choosing this entry will ask the user to select a (secondary) VMR, which will be loaded and its data stored in the currently selected VMR'sRunTimeVars.UndoBufferproperty, allowing to draw data from a different object into the currently selected VMR's data (using the “undo-drawing” functionality) * Export to Nifti - exports the current VMR object as a Nifti file without reslicing ===== VTC menu ===== This menu is only visible if the currently selected slicing (voxel-based anatomical) object is of type VTC. * Mask with VMR - if more than one VMR is open, a small listbox dialog (listdlg) showing the currently loaded VMRs will be shown, and the currently selected VTC will then be masked so that voxel timecourses where the VMR data is below visible threshold will be set to all-zeros * Mean/Standard dev. maps - creates a VMP with two maps, one representing the mean (over time) and the other representing the standard deviation; this can be used for quick diagnostics (homogeneity as well as spatially isolated temporal outliers) * Export to Nifti - exports the current VTC object as a Nifti file without reslicing ===== GLM menu ===== This menu is only visible if the currently selected slicing (voxel-based) statistical object is of type GLM. * Beta/scatter plot - a separate figure (GLM betaplot UI) will be opened, showing either bar graphs (incl. SD/SE indicators) or a scatter plot (for which RFX covariates must be already pre-configured); for additional information, please follow the previous link * Generate MDM… - using the references in the GLM object, an MDM is created, for which a file saving dialog is then presented; this allows to re-compute statistics with different settings in case the original MDM is not available * Mask with VMR - applies the currently selected VMR (and thus only works if the selected slicing object is of type VMR!) as a mask to the GLM in a way that, for all maps in the GLM object, voxels where the VMR data is below the visibility threshold will be set to 0 * Export to Nifti - exports the current GLM object as a Nifti file without reslicing ===== VMP menu ===== This menu is only visible if the currently selected slicing (voxel-based) statistical object is of type VMP. * Combine VMPs… - if more than one VMP (with the same spatial layout, defined by resolution, offset and size of maps) is loaded, this will present a listbox dialog (listdlg) showing the other loaded VMP objects, from which one or several objects can be selected; subsequently all maps found in the selected objects will be added at the end of the currently selected VMP * Create MSK… - for the currently selectedMapsa MSK object will be created that can then be used during subsequent GLM regressions; if several maps are selected, the user will be asked to specify how these maps are to be combined (intersection, union, or average) * Create SMP… - if a surface object is loaded (and selected), the currently selected VMP will be sampled at the positions of the vertices of the selected surfaces and this data will be stored in appropriate SMP objects; a file saving dialog (uiputfile) will be shown for the newly created object so as to simplify pairing SMPs with their according SRF object * Export maps to Analyze - this will save all currently selectedMapsfrom the selected VMP into Analyze (HDR format) files * Mask with VMR - applies the currently selected VMR (and thus only works if the selected slicing object is of type VMR!) as a mask to theMapsin the VMP, so that voxels where the VMR data is below the visibility threshold will be set to 0 * Mask with VOI clusters - applies the currently selected clusters from the list of all clusters (VOIs) as a mask, so that all voxels that are not covered by any of the selected clusters will be set to 0 * Export to Nifti - exports the current VMP object as a Nifti file without reslicing ===== SRF menu ===== This menu is only visible if an object of type SRF is loaded. * Reload from disk - reloads the currently selected surface file from disk (only functional if the object has been loaded or saved before and a filename is associated with it) * Smooth… - brings up a dialog with smoothing settings (number of iterations, smoothing force) followed by passing these settings to a call to SRF::Smooth ===== HDR menu ===== This menu is only visible if the currently selected slicing (voxel-based anatomical) object is of type HDR/NII. * Toggle oblique slicing - if selected, this will set (or revert) the spatial vectors (data extension directions) of the currently selected HDR object to be oblique with the main coordinate axes (L/R, A/P, S/I) ===== Statistics menu ===== This menu is only visible when the GUI is in minimized mode! * Close OBJ - closes (unloads) the currently selected voxel-based statistical object (if the object was created by the GUI during the current session, the object will also be cleared and its memory freed) * Map properties - invokes the map properties edit dialog (only visible when exactly one map of a VMP object is selected) * Compute formula - invokes the AFT::ComputeFormula interface (only visible when a HDR/HEAD/VMP object is selected) * Switch to stats object - a sub-menu populated with a list of currently loaded voxel-based statistics objects * Select map - a (list of) sub-menu(s) populated with a list of the map names of the currently selected voxel-based statistics object (with selected maps being checked) * Multi-select maps - opens a listbox dialog that allows to select multiple maps * Select CONDITION for all subjects - allows to copy map selection of one subject across subjects (only visible when a RFX-GLM object is selected) * Delete selected maps - deletes the selected maps ===== Analysis menu ===== * Compute multi-study GLM - this will open up a dialog that can be used to interactively configure (or load and alter) an MDM object, followed by running the required regression (RFX only at this time); for further reading, please visit the Compute multi-study GLM UI page, as well as the MDM::ComputeGLM reference page * Contrast manager - this will open up the (for now RFX-only) contrast manager, which requires at least one RFX-GLM to be loaded; if several RFX-GLMs are loaded, the user can choose between these in the Contrast Manager UI, and if a GLM is selected as current statistical object, this GLM will be the pre-selection * Single-level RFX mediation - this will open up the RFX mediation UI (which requires at least one RFX-GLM to be loaded; if several RFX-GLMs are loaded, the user can choose between these in the UI) * ECG Heart-rate data analysis - this will open up the ECG/GSR data analysis UI ===== Visualization menu ===== * Create montage images - this will open up the Image Montage UI * Create rendered images - this will open up the Rendering UI * Create VOI condition plots - this function requires clusters (VOIs) to be defined in the main UI window, for which functional data will be extracted and averaged according to conditions; first, the user will be asked to select an MDM file (which must be PRT-based!), after which the protocol (PRT) files will be read and the list of conditions to be available in subsequent plots will be presented in a listbox dialog (listdlg) for selection; following data extraction and computation, the MDM VOI condition average plot window will open * Reload VOI condition plots - as part of the MDM VOI condition average UI, once extracted data can be saved into a MAT file, which can then be reloaded at a later time (without having to re-extract and possibly regress the data), which is usually much more time efficient ===== VOI menu ===== This menu is only visible when the GUI is in minimized mode! * Go to nearest cluster (peak) - sets the cursor position to the nearest peak in the list of VOIs (first voxel for each VOI) * Select cluster… - a (list of) sub-menu(s) populated with the names of the VOIs found in the cluster table list ===== Tools menu ===== * SPM → BV conversion… - this is a sub-item menu with … * SPM.mat → PRT(s)… - a function that allows the user to select oneSPM.matfile, which will be read and for each session (.Sess) found therein, a protocol (PRT file) will be written to disk * SPM.mat → SDM(s)… - likewise, this function allows the user to select oneSPM.matfile, and for each session one SDM file will be written to disk * alphasim… - if the currently selected (voxel-based) statistical object is a VMP, this will first create a mask from all non-zero voxels of the currently selected map(s) and then request the user to enter a number of iterations, after which thealphasimfunction is called and results are being reported in the Matlab console window * Rename DICOM files… - this will call renamedicom without any parameters, which will open its own UI * SPM5 preprocessing… - this will open the UI of spm5_preprojobs, a function that configures a standardized SPM5/SPM8-based stream of preprocessing steps * tdclient… - this opens up the neuroelf_gui - tdclient UI * Quality assessment - this is a sub-item menu with … * Assess fMRI quality… - a quality assessment of a selected (list of) files, using the fmriquality and fmriqasheet functions with a predefined set of options * Visualize SPM motion params… - this brings up a file selector and then passes the selected files to the showspmrparams function ===== Help menu ===== * About NeuroElf GUI…** - next to some thanks and references, the information shown contains a list of keyboard shortcuts available in the NeuroElf GUI
neuroelf_gui_-_menu_tree.1304798342.txt.gz · Last modified: 2011/05/07 19:59 by jochen
