neuroelf_gui_-_mkda_ui
Table of Contents
NeuroElf - Multi-level Kernel Density Analysis (MKDA) UI
Motivation
When compiling a list of coordinates to be considered for an MKDA, it is often cumbersome to configure the various analyses that can be run (e.g. eliminating studies/contrasts from the analysis that do not comply with a certain methodology such as having used standardized stimuli). To overcome this relatively difficult task, NeuroElf comes with a UI that tries to simplify this as much as possible.
Requirements
Please read the more conceptually oriented MKDA article.
GUI layout
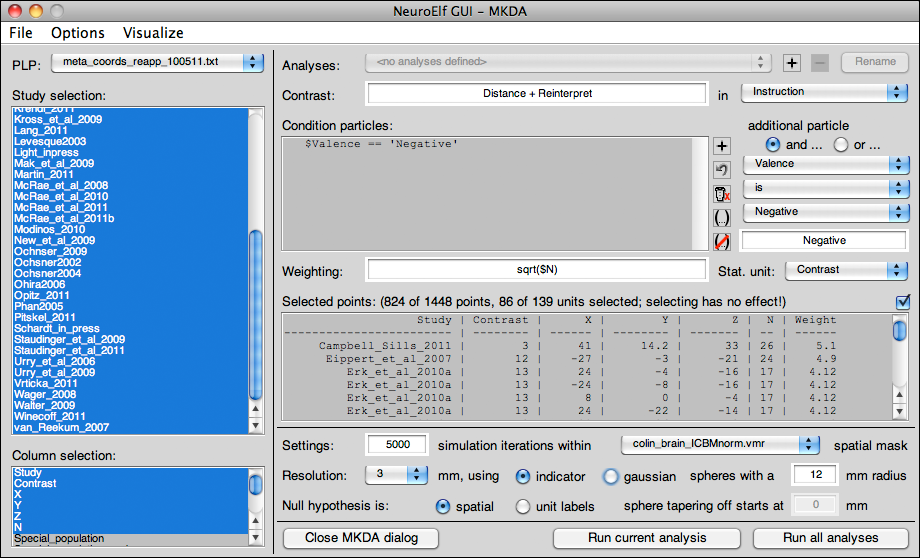
The MKDA UI is available via the Meta Analysis entry of the Analysis menu. When called, the following dialog pops up:
Controls
The following controls are available:
- PLP selector (dropdown): if more than one PLP object is loaded, you can switch between those (e.g. in case you have several revisions of the same database)
- Study selection (listbox): by (de-) selecting studies, you can re-run the same analyses discarding points from some studies without having to create several versions of the database
- Column selection (listbox): the selection of columns will alter the output of the listed points (listbox, see below)
- Analyses (dropdown): this control stores all configured analyses, which allows to easily define several contrasts and/or settings without having to create separate database files (PLP objects)
- Adding a new analysis (+ button): upon clicking this button, NeuroElf asks for an analysis name, and then duplicates the currently configured settings and makes them available for further editing
- Removing the current analysis (- button): if several analyses are configured, using this button allows to delete one of them (e.g. for contrasts/configurations that yielded inconclusive results)
- Renaming the current analysis: clicking this button will ask for a new name for the currently selected analysis
- Contrast (edit field): you can use this field to enter a contrast made up of the contrast terms that are listed automatically when selecting a column name containing contrast particles; a differential contrast can be entered using the greater than (>) sign between particles (e.g.
item1 + item2 > item3 + item4) - Contrast column (dropdown): use this control to select from which column in the database the contrast particles are selected (if you wish to combine several factors for contrasts, you MUST configure them in a compound column while creating the database file!); upon selection, the contrast edit field will be set to a list of all unique terms found in this column
- Condition particles (listbox): this is a text-based representation of the conditions which are used to sub-select coordinates from the database for each configured analysis; just as with Matlab expressions, conditions can be made up of several conditional statements (particles) conjoined with either AND or OR operators and evaluated in any desired precedence using parentheses around sets of particles; to configure the condition particles use the subsequent controls!
- Adding a condition particle (+ button): use this button to add the condition as configured on the right side of the button to the list of particles; if any particles are already in the list, the operator is placed in front of the newly added particle
- Setting the current particle (arrow button):
Usage notes
The following steps should be performed for any analysis:
- loading or importing a PLP object
- if you have already imported a PLP at a previous session, use the
Load PLPentry from the File menu - otherwise using the
Import PLPentry from the File menu; it is highly recommend to save any imported object as a PLP, as this will then automatically store any configured contrast, conditions, and settings alongside the PLP file
- either selecting or creating an analysis
- in case you have already created an analysis at an earlier time, you can simply select an analysis; if you wish to create multiple analysis with similar settings, it is recommended to first create one of these and then use the “Add new analysis” button to duplicate the settings and retouch the settings as desired
- otherwise, start by choosing a contrast column, then configure the contrast terms (particles), followed by setting up any conditional statement you would like to use for further sub-selection of coordinates from the database
- configure all further settings (spatial layout, etc.)
- run the current analysis (or all configured analyses), which will create one VMP per analysis (and possibly one additional VMP with just the resulting summary maps)
Outcome description
neuroelf_gui_-_mkda_ui.txt · Last modified: 2011/12/08 22:56 by jochen